*The picture in the header was taken from here.
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
______________________________________________________________________________________________________
Protein Interactions
By looking at protein interactions, we will be able to identify the proteins that the UBE3A protein interacts with. This information can give some clues on how the UBE3A protein interacts with other proteins, what other functions the UBE3A protein is involved in and how the missing or defective protein can give rise to Angelman Syndrome.
Here, we used 2 protein interactions databases, STRING 9.0 and MINT to generate the UBE3A protein interaction network.
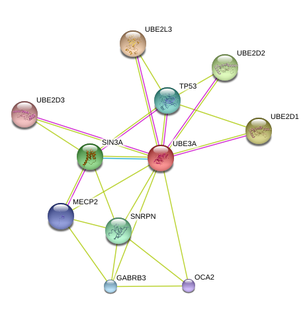
STRING 9.0
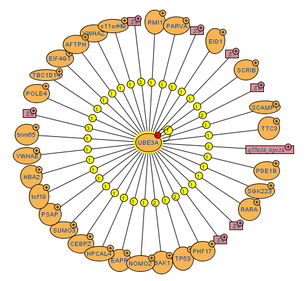
MINT
Analysis
The proteins listed by STRING 9.0 made sense when looking at the gene ontology of UBE3A. UBE2L3, UBE2D1, UBE2D2, UBE2D3 are all ubiquitin-conjugating enzyme [2] and they assist UBE3A in attaching ubiquitin to other proteins. As discussed in the Protein Domain section, the ubiquitin-conjugating enzyme transfers ubiquitin in the form of a thioester to ubiquitin-protein ligase before the ubiquitin-protein ligase transfers the ubiquitin to targeted substrates. TP53 is a tumor suppressor protein and involves in growth arrest or apoptosis [2]. This suggests that TP53 might play a part in neuronal apoptosis. According to GeneCards, SNRPN plays a role in maternal imprinting and this is consistent with the current knowledge that in the brain, only the maternal copy of UBE3A is expressed due to genomic imprinting. GABRB3 is the major inhibitory neurotransmitter in the brain [2] and it could be possible that UBE3A interacts with it to regulate neuronal activities in the brain. UniProt reports that patients with defective GABRB3 show signs of chronic insomnia and epilepsy. These symptoms are also manifested by AS patients so it is suggestive that the missing functional UBE3A affects the expression of GABRB3, which gives rise to the 2 mentioned symptoms.
Besides using STRING 9.0 and MINT, a review paper by Mabb et al. was also referred to. Mabb et al. explained how the UBE3A protein interacts with other proteins and how this could possibly contribute to neuronal morphology and neural circuits’ development. During synaptogenesis, UBE3A attaches ubiquitin to RhoA guanine nucleotide exchange factor (RhoA-GEF) Ephexin-5 and marks it for degradation by the ubiquitin proteasome system (UPS). This inactivates RhoA and assists the formation of dendritic spines.
Besides that, UBE3A also ubiquitinates and promotes the degradation of Arc (activity-regulated cyto-skeleton-associated protein). Arc is only found in the brain and is rapidly upregulated in response to increases in neuronal activity. UBE3A regulates AMPAR (AMPA receptor) endocytosis by controlling the amount of Arc. By doing so, UBE3A helps to restructure pre-existing synapses and this allows the development of functional neural circuits.
In cases where there is excessive Arc, endocytosis of GluA1-containing AMPARs from glutamatergic synaptic sites is unusually high. This leads to a reduction in excitatory synaptic transmission and also an increase in the number of silent (AMPAR-lacking) synapses. The defective synapses and neural circuits could be the possible cause to the various Angelman Syndrome phenotypes, such as intellectual and developmental delay, and impaired social skills.
Mabb et al.also mentioned that the UBE3A protein interacts with the tumor suppressor protein p27. In the cortex, p27 protein promotes neuronal differentiation and migration in cortical projection neurons. With the absence or low amount of the UBE3A protein, there is an increase in the p27 protein level, which might cause the premature migration and differentiation of cortical neuronal progenitors, and also change the laminar architecture of the cortex.
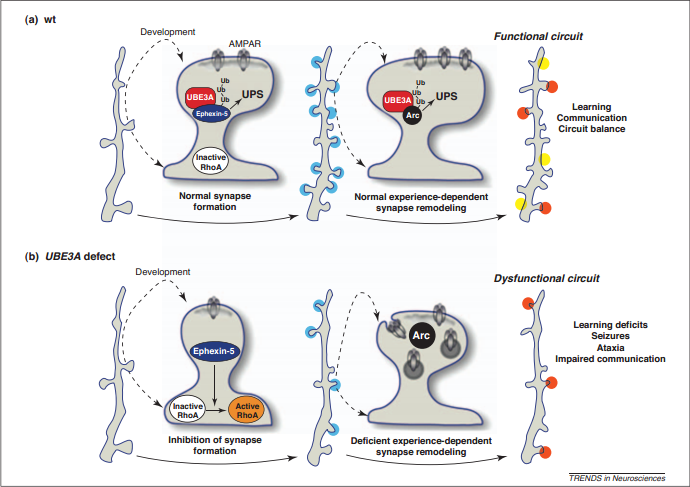
Figure 3: A diagram illustrating how UBE3A interacts with other proteins and how this possibly contributes to neuronal morphology and neural circuits’ development. Diagram taken from [4].
References:
[1] GeneCards
[2] STRING 9.0
[3] MINT
[4] UniProt
[5] Mabb, A. M., Judson, M. C., Zylka, M. J., Philpot, B. D. (2011). Angelman syndrome: insights into genomic imprinting and neurodevelopmental phenotypes. Trends in Neurosciences, 34(6): 293-303. doi: 10.1016/j.tins.2011.04.001
[1] GeneCards
[2] STRING 9.0
[3] MINT
[4] UniProt
[5] Mabb, A. M., Judson, M. C., Zylka, M. J., Philpot, B. D. (2011). Angelman syndrome: insights into genomic imprinting and neurodevelopmental phenotypes. Trends in Neurosciences, 34(6): 293-303. doi: 10.1016/j.tins.2011.04.001
If you find my website helpful, please consider donating to the Foundation for Angelman Syndrome Therapeutics (FAST)

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677