*The picture in the header was taken from here.
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
______________________________________________________________________________________________________
DNA Motifs
DNA has short, recurring sequences, known as DNA motifs [4]. In general, they are presumed to have a biological function, such as binding sites for nucleases and transcription factors [4]. By looking at the sequence motifs, we could deduce the function of the gene or how the gene interacts with other genes.
MOTIF
MOTIF was used to identify the DNA motifs of the UBE3A gene. Below are the 11 identified motifs:
|
Motif Name: EGF_1
Prosite ID: PS00022 Description: EGF-like domain signature 1 Pattern: C-x-C-x(2)-{V}-x(2)-G-{C}-x-C. Motif Name: CTCK_1 Prosite ID: PS01185 Description: C-terminal cystine knot signature. Pattern: C-C-x(13)-C-x(2)-[GN]-x(12)-C-x-C-x(2,4)-C. Motif Name: IGFBP_N_1 Prosite ID: PS00222 Description: Insulin-like growth factor-binding protein (IGFBP) N-terminal domain signature. Pattern: [GP]-C-[GSET]-[CE]-[CA]-x(2)-C-[ALP]-x(6)-C. Motif Name: TUBULIN Prosite ID: PS00227 Description: Tubulin subunits alpha, beta, and gamma signature. Pattern: [SAG]-G-G-T-G-[SA]-G. Motif Name: 4FE4S_FER_1 PrositeID: PS00198 Description: 4Fe-4S ferredoxin-type iron-sulfur binding region signature. Pattern: C-x-{P}-C-{C}-x-C-{CP}-x-{C}-C-[PEG]. Motif Name: DEFENSIN PrositeID: PS00269 Description: Mammalian defensins signature. Pattern: C-x-C-x(3,5)-C-x(7)-G-x-C-x(9)-C-C. |
Motif Name: INTEGRIN_BETA
Prosite ID: PS00243 Description: Integrins beta chain cysteine-rich domain signature. Pattern: C-x-[GNQ]-x(1,3)-G-x-C-x-C-x(2)-C-x-C. Motif Name: ANAPHYLATOXIN_1 Prosite ID: PS01177 Description: Anaphylatoxin domain signature. Pattern: [CSH]-C-x(2)-[GAP]-x(7,8)-[GASTDEQR]-C-[GASTDEQL]- x(3,9)-[GASTDEQN]-x(2)-[CE]-x(6,7)-C-C. Motif Name: THIOLASE_3 Prosite ID: PS00099 Description: Thiolases active site. Pattern: [AG]-[LIVMA]-[STAGCLIVM]-[STAG]-[LIVMA]-C-{Q}-[AG]- x-[AG]-x-[AG]-x-[SAG]. Motif Name: VWFC_1 Prosite ID: PS01208 Description: VWFC domain signature. Pattern: C-x(2,3)-C-{CG}-C-x(6,14)-C-x(3,4)-C-x(2,10)-C-x(9,16) -C-C-x(2,4)-C. Motif Name: 2FE2S_FER_1 PrositeID: PS00197 Description: 2Fe-2S ferredoxin-type iron-sulfur binding region signature. Pattern: C-{C}-{C}-[GA]-{C}-C-[GAST]-{CPDEKRHFYW}-C. |
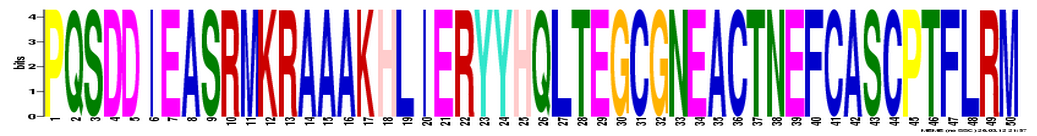
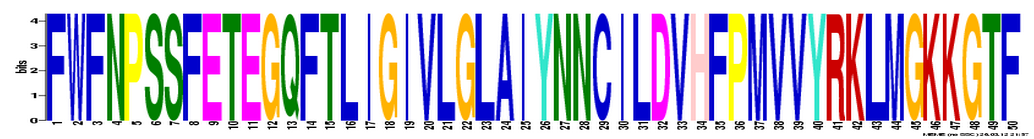
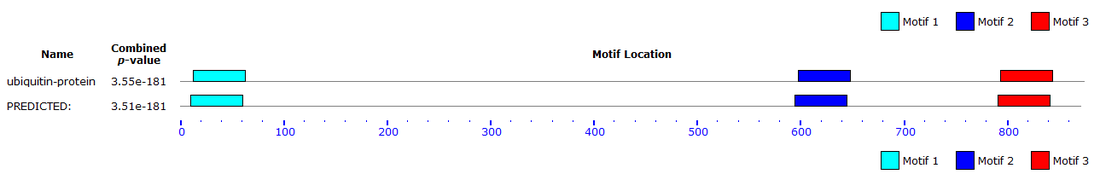
MEME
MEME restricted the length of a gene sequence that can be inputted to 60,000bp. The UBE3A gene is 101780bp long so the full gene sequence was not used. All the homologous protein sequences were used instead. The results can be viewed here.
Analysis
2 databases were used to identify the DNA motifs: MOTIF and MEME. MOTIF found 11 motifs while MEME (search done by using homologous protein sequences) identified 3 motifs. INTEGRIN_BETA and CTCK_1 are worth mentioning because they involve the cysteine amino acid. According to Pfam, the active site of the UBE3A protein is a cysteine. Therefore, these motifs might contribute to the active site of the UBE3A protein.
References:
[1] MOTIF
[2] MEME
[3] Pfam
[4] D'haeseleer, P. (2006). What are DNA sequence motifs? Nature Biotechnology, 24(4): 423 - 425. doi:10.1038/nbt0406-423
[5] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
If you find my website helpful, please consider donating to the Foundation for Angelman Syndrome Therapeutics (FAST)

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677