*The picture in the header was taken from here.
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
______________________________________________________________________________________________________
Gene Ontology
Gene ontology refers to the properties of shared biological elements (such as genes and proteins) across species. There are 3 domains, namely: biological process, cellular component, and molecular function. By looking at gene ontology, inference on how the UBE3A gene affects Angelman Syndrome patients can be made.
UBE3A gene ontology information is obtained via AMIGO. AMIGO returned a list of result but only those that are considered relevant to Angelman Syndrome are listed here. Click on each result for more details.
Biological Process:
- Androgen Receptor Signaling Pathway
- Brain Development
- Positive Regulation of Phosphatidylinositol 3-kinase Cascade
- Positive Regulation of Transcription From RNA Polymerase II Promoter
- Protein K48-linked Ubiquitination
- Protein Modification Process
- Ubiquitin-dependent Protein Catabolic Process
Cellular Component:
Molecular Function:
- Amino Acid Ligase Activity
- Protein Binding
- Transcription Coactivator Activity
- Ubiquitin-protein Ligase Activity
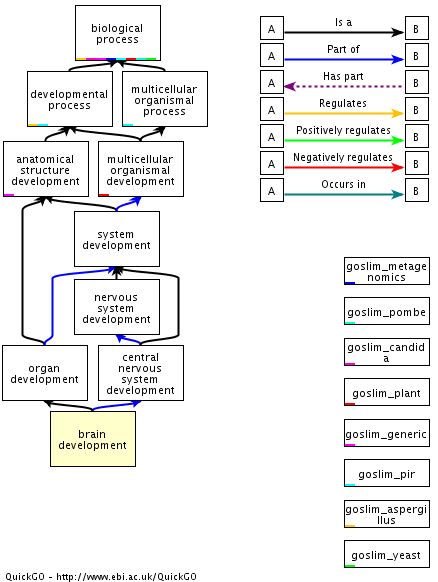
Figure 1: This graphical view shows how the UBE3A gene involves in the brain development pathway. Diagram taken from AMIGO
Analysis
AMIGO shows that the UBE3A gene is involved in protein modification and this is most likely by ubiquitinating other proteins. By attaching ubiquitin to proteins, the proteins are marked for degradation by the ubiquitin proteasome system (UPS) [2]. This explains why UBE3A is dependent on ubiquitin for catabolic process, as identified by AMIGO.
AMIGO identified that the UBE3A products are found in the proteasome complex. This makes sense as the E6AP ubiquitin-protein ligase encoded by UBE3A acts with proteases to mark and degrade proteins. E6AP ubiquitin-protein ligase is also found in the nucleus, which is consistent with the data obtained from Dindot et al.'s experiment [3].
References:
[1] Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. (2000). Gene Ontology: Tool for the unification of biology. Nature Genetics, 25(1): 25–29. doi:10.1038/75556
[2] Mabb, A. M., Judson, M. C., Zylka, M. J., Philpot, B. D. (2011). Angelman syndrome: insights into genomic imprinting and neurodevelopmental phenotypes. Trends in Neurosciences, 34(6): 293-303. doi: 10.1016/j.tins.2011.04.001
[3] Dindot, S. V., Antalffy, B. A., Bhattacharjee, M. B., Beaudet, A. L. (2007). The Angelman syndrome ubiquitin ligase localizes to the synapse and nucleus, and maternal deficiency results in abnormal dendritic spine morphology. Human Molecular Genetics, 17(1): 111-118. doi:10.1093/hmg/ddm288
If you find my website helpful, please consider donating to the Foundation for Angelman Syndrome Therapeutics (FAST)

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677
