*The picture in the header was taken from here.
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
______________________________________________________________________________________________________
Protein Domain
A protein domain is a part of a protein sequence that folds independently to form a stable 3D structure and can usually function independently from the rest of the protein chain [1]. Proteins can have multiple domains, thereby, conferring them multiple functions. By looking at the domains that a protein has, we can infer what the functions of the protein are and how it interacts with other proteins or genes.
3 protein domain databases, namely Pfam, SMART, and PROSITE were used to identify the protein domains in the Ubiquitin-protein ligase E3A Isoform II (E6AP) protein.
Pfam
Pfam found one domain in the E6AP protein, the HECT domain (Homologous to E6-AP Carboxyl Terminus or also known as ubiquitin transferase), with an e-value of 4.6e-93. Pfam suggested that the location of the HECT domain to be from 577 to 874. Besides that, Pfam also located the active site of the domain to be at 843, as indicated by the purple diamond in Figure 1.
Pfam found one domain in the E6AP protein, the HECT domain (Homologous to E6-AP Carboxyl Terminus or also known as ubiquitin transferase), with an e-value of 4.6e-93. Pfam suggested that the location of the HECT domain to be from 577 to 874. Besides that, Pfam also located the active site of the domain to be at 843, as indicated by the purple diamond in Figure 1.

Figure 1: The green box shows the location of the HECT domain while the purple diamond indicates the active site. Diagram taken from Pfam
SMART
SMART was later used, in hope of searching for more domains but only the HECT domain was found (with a slightly more significant e-value, 4.72e-180). SMART showed that the location of the domain to be from 545 to 875, a little ahead than that found by Pfam.
SMART did not show the location of the active site but showed a coiled coil region from 159 to 187 (indicated by the green box) and a low complexity region (LCR) at position 394-406, as represented by the pink box. According to SMART, coiled coils refer to groups of long alpha-helices that coil together[3]. Coletta et al. explains that LCR is a region of protein sequences with biased amino acid composition[4].
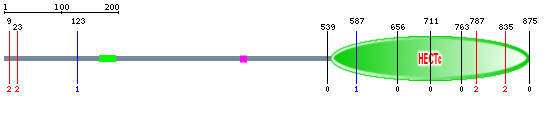
Figure 2: The big green oval shape represents the HECT domain, the small green box shows the coiled coil region and the pink box indicates the low complexity region. Diagram taken from SMART
Structure of HECT Domain of E6AP
In 1999, Huang, et al. have determined the crystal structure of the human E6AP HECT domain when bound to the human UbcH7 ubiquitin-conjugating enzyme [5]. The crystal structure revealed that the E6AP hect domain NH2-terminal lobe consisted of 12 α helices and six β strands while the COOH-terminal lobe comprised of six α helices and four β strands [5]. The structure is shown in Figure 3. The long cylinder structures are the α helices while the arrow-like structures are the β strands. The yellow structure represents the active site.
Do note that the crystal structure is for residues from 495 to 852. However, the 3 protein domains database (Pfam, SMART and PROSITE) suggest that the HECT domain is located at ~545-875.
In 1999, Huang, et al. have determined the crystal structure of the human E6AP HECT domain when bound to the human UbcH7 ubiquitin-conjugating enzyme [5]. The crystal structure revealed that the E6AP hect domain NH2-terminal lobe consisted of 12 α helices and six β strands while the COOH-terminal lobe comprised of six α helices and four β strands [5]. The structure is shown in Figure 3. The long cylinder structures are the α helices while the arrow-like structures are the β strands. The yellow structure represents the active site.
Do note that the crystal structure is for residues from 495 to 852. However, the 3 protein domains database (Pfam, SMART and PROSITE) suggest that the HECT domain is located at ~545-875.
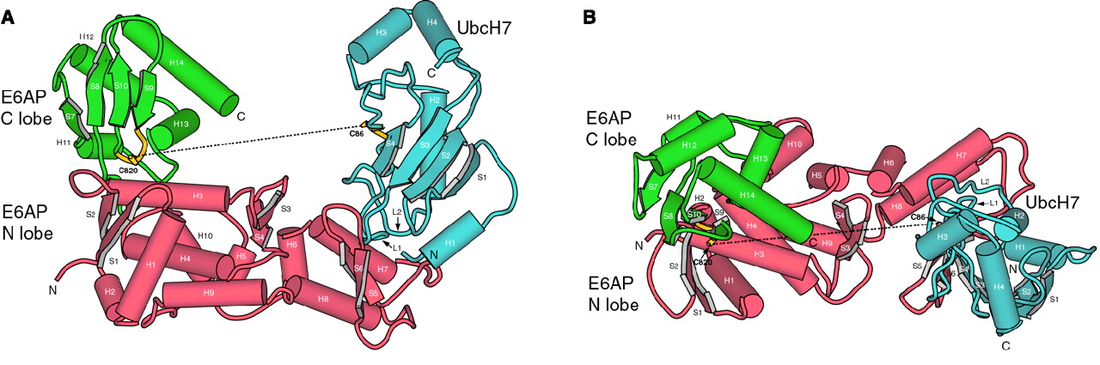
Figure 3: The crystal structure of the E6AP HECT domain by Huang, L. et al.
Diagram A shows the U-shaped structure formed by the E6AP HECT domain-UbcH7 complex while Diagram B is the orthogonal view of the overall structure of the complex.
Diagram A shows the U-shaped structure formed by the E6AP HECT domain-UbcH7 complex while Diagram B is the orthogonal view of the overall structure of the complex.
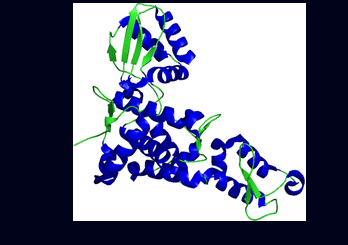
Figure 4: The structure of E6AP HECT domain based on Huang et al's crystal structure of the E6AP HECT domain-UbcH7 complex .
Click here for an animated version of the structure. Diagram taken from The Pawson Lab
Click here for an animated version of the structure. Diagram taken from The Pawson Lab
Analysis
It is interesting to see there is only one protein domain in the Ubiquitin-protein ligase E3A Isoform II or E6AP protein. This domain is the HECT domain, short for Homologous to E6-AP Carboxyl Terminus. All 3 protein domains databases did not give the same starting location for this domain but they identified the same ending location and the same location for the active site. The slight discrepancy between the starting locations is probably due to the different algorithm that the databases used to determine the location of the domain.
The active site is identified to be a cysteine at position 843. According to Pfam, ubiquitin-protein ligase accepts ubiquitin from an E2 ubiquitin-conjugating enzyme in the form of a thioester, and later transfers the ubiquitin directly to targeted protein molecules. The cysteine residue at the active site comes to play during the ubiquitin-thiolester formation.
Huang, et al. have identified that mutation of Thr(819), Asn(822), or Phe(821) in the active site loop to Ala affected the ability of the HECT domain to form the ubiquitin thioester intermediate by about 70%. The Arg(506) , Glu(539), and Glu(550) residues on the N-lobe form a solvent-exposed salt-bridge network adjacent to the catalytic cysteine and to Asp(607). It was observed that if there was mutation in any one of these four conserved residues on the N lobe, the ubiquitin-thioester formation was reduced by more than 90%. This suggested that the N-lobe portion of the cleft was needed to form the ubiquitin thioester intermediate.
In the same paper, it was mentioned that previous studies had shown that deletion of the last six residues of E6AP had caused the elimination of isopeptide bond formation between ubiquitin and the substrate protein without substantially affecting the formation of the ubiquitin-thioester intermediate. This might imply that the residues in the C lobe are important for the catalysis of isopeptide bond formation, instead of ubiquitin thioester intermediate formation.
*The number in parenthesis refers to the position of the amino acid.
References:
[1] Richardson J. S. (1981). The anatomy and taxonomy of protein structure. Advances in Protein Chemistry, 34:167-339. doi:10.1016/S0065-3233(08)60520-3
[2] Pfam
[3] SMART
[4] SMART Glossary
[5] Coletta, A., Pinney, J. W., Solís, D. YW., Marsh, J., Pettifer, S. R., Attwood, T. K. (2010). Low-complexity regions within protein sequences have position-dependent roles. BMC Systems Biology, 4:43. doi:10.1186/1752-0509-4-43
[1] Richardson J. S. (1981). The anatomy and taxonomy of protein structure. Advances in Protein Chemistry, 34:167-339. doi:10.1016/S0065-3233(08)60520-3
[2] Pfam
[3] SMART
[4] SMART Glossary
[5] Coletta, A., Pinney, J. W., Solís, D. YW., Marsh, J., Pettifer, S. R., Attwood, T. K. (2010). Low-complexity regions within protein sequences have position-dependent roles. BMC Systems Biology, 4:43. doi:10.1186/1752-0509-4-43
If you find my website helpful, please consider donating to the Foundation for Angelman Syndrome Therapeutics (FAST)

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677

Created by Jonathan Mok
[email protected]
Last updated 02/23/2012
Genetics 677
